Cancer Genomics Cloud presented at AACR
The Cancer Genomics Cloud at AACR
On Sunday, Dr Brandi Davis-Dusenbery presented the Seven Bridges Cancer Genomics Cloud (CGC) to a packed room at the American Association of Cancer Research (AACR) annual meeting in New Orleans. We are thrilled to see so much enthusiasm about the cloud pilots, and that so many people were keen to discuss technical details of the CGC!
This presentation was part of a session dedicated to the NCI Cancer Genomics Cloud Pilots, which are intended to explore innovative methods for accessing and computing on large genomic data, and to facilitate access to The Cancer Genome Atlas (TCGA). Our colleagues at the Institute of Systems Biology and the Broad Institute also presented status updates on their own cloud pilot projects.
TCGA data in the cloud
Brandi showed how the CGC enables users to access more than a petabyte of multidimensional data from TCGA, and more importantly to explore the dataset using visualization and powerful metadata filters.
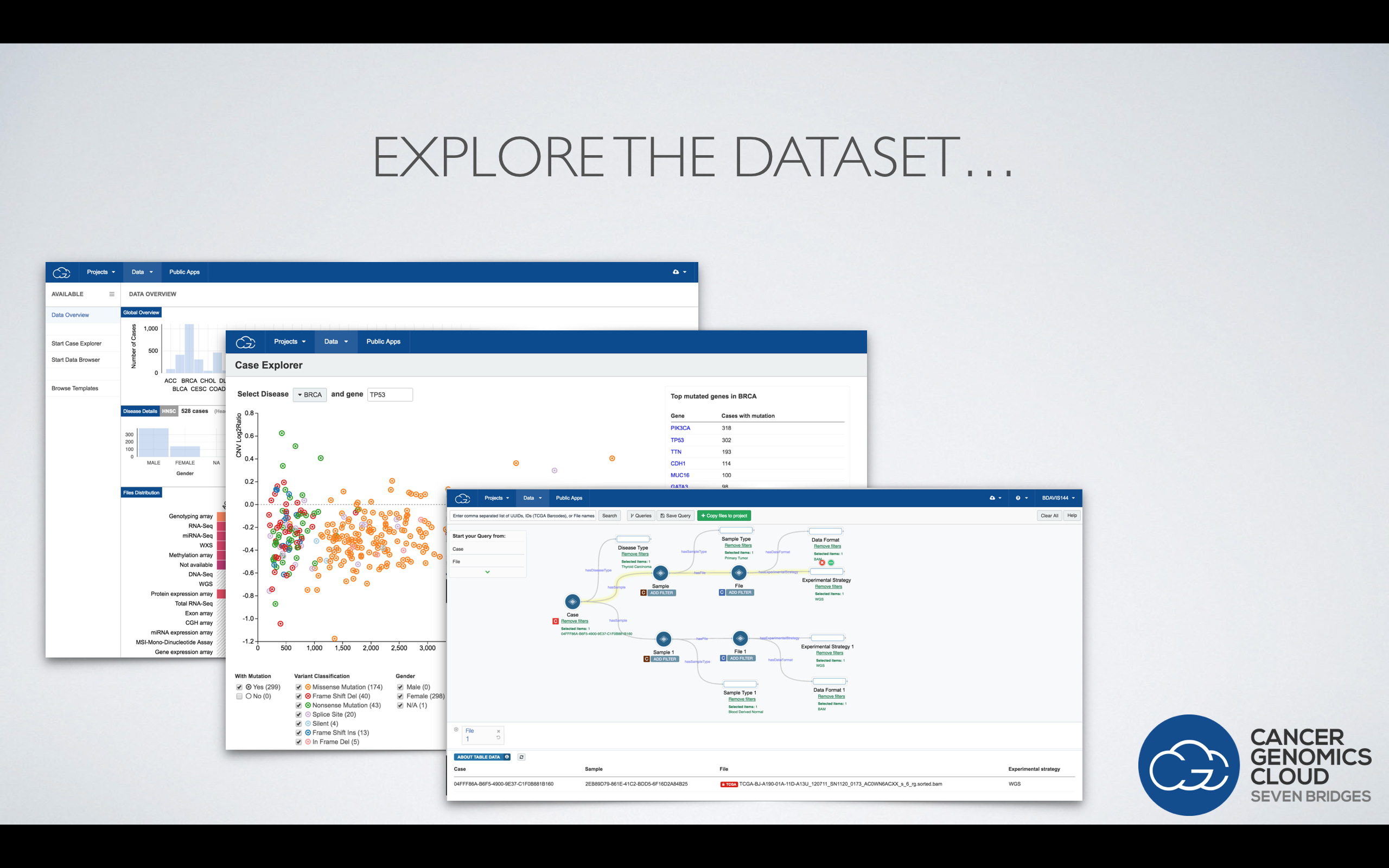
Users can then immediately run an analysis of the data on the cloud in a secure, collaborative environment. All analyses done on the CGC are fully reproducible and rememberable. Users can bring their own data and tools to the CGC, or use those already on the platform.
The CGC launched in February 2016, and so far nearly 500 researchers have signed up to use its resources.
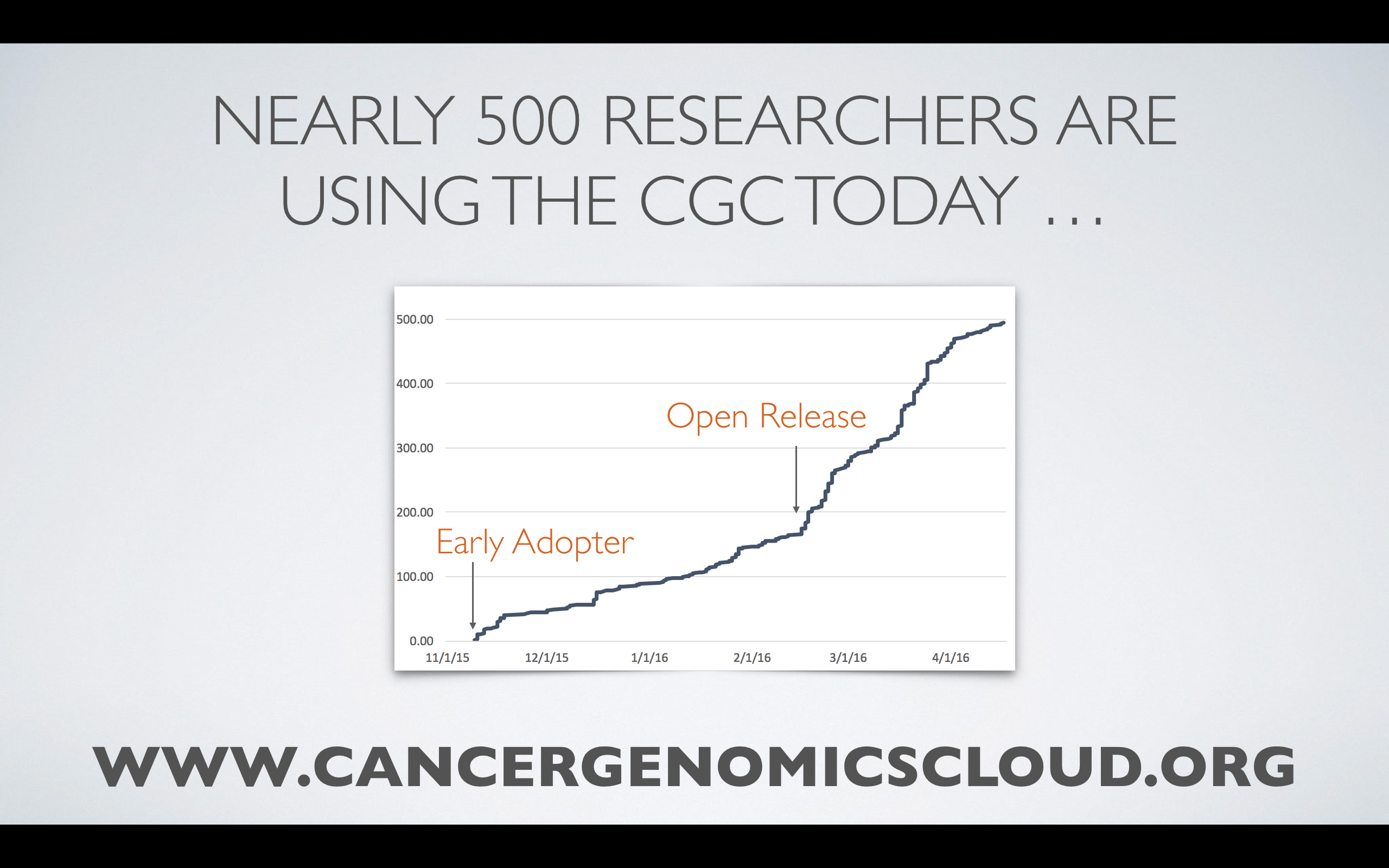
Cancer researchers, including graduate students and postdoctoral researchers, are encouraged to sign up and explore TCGA data and available tools. The CGC is available to all, and over $1,000,000 in credits is available to users to analyze and store data.
CGC in action
In the same session Jeffrey Chuang, a CGC user from the Jackson Laboratory for Genomic Medicine presented results of research into the effects of tumor heterogeneity on survival across tumor types.
Jeff Chuang of @jacksonlab. His analyses of TNBC breast #cancer xenograft data -> tumor evolution. #AACR16 pic.twitter.com/bXCp0NA9C2
— Mark Wanner (@markgenome) April 18, 2016
His team used the CGC to analyze 1,741 tumor–normal pairs across multiple tumor types using different variant callers. The data were extracted by filtering the TCGA dataset using the CGC—all done in the cloud without needing to download the massive TCGA dataset. The analysis, which comprised 5,275 tasks on almost 3,500 BAM files, completed in just 3 days.
We think this neatly illustrates the power of the CGC to rapidly access and analyze the massive TCGA dataset.
NCI Initiatives
Discussion of the cloud pilots was one of several NCI/NIH-sponsored sessions, which showcased the NCI initiatives aimed at understanding the biology of cancer. Among the highlights:
- Jean Zenklusen gave an update on the TCGA project, which has provided a rich dataset for understanding cancer biology and is scheduled to complete this year. He argued that TCGA is the world’s best example of ‘team science’ in biology. On completion, focus will shift to other initiatives that center on understanding the relationship of cancer genomics to clinical responses.
- Zhining Wang spoke on the Genomic Data Commons (GDC), which will launch later this year with the aim of providing the cancer research community with a unified data repository. The GDC aims to allow researchers to upload their own data, which will undergo quality control and be harmonized so that data can be compared across disease types. Evaluation of the NCI cloud pilots will inform how the GDC data are made accessible to researchers.
- We also heard about clinical initiatives from the NCI Center for Cancer Genomics, including the Exceptional Responders Initiative that aims to find molecular indicators in patients who respond exceptionally well to treatment; and ALCHEMIST, which studies whether treatment based on genotype improves cure rates in non-small cell lung cancer.